Phosphorylation Networks and Cellular Signaling | |
|
|
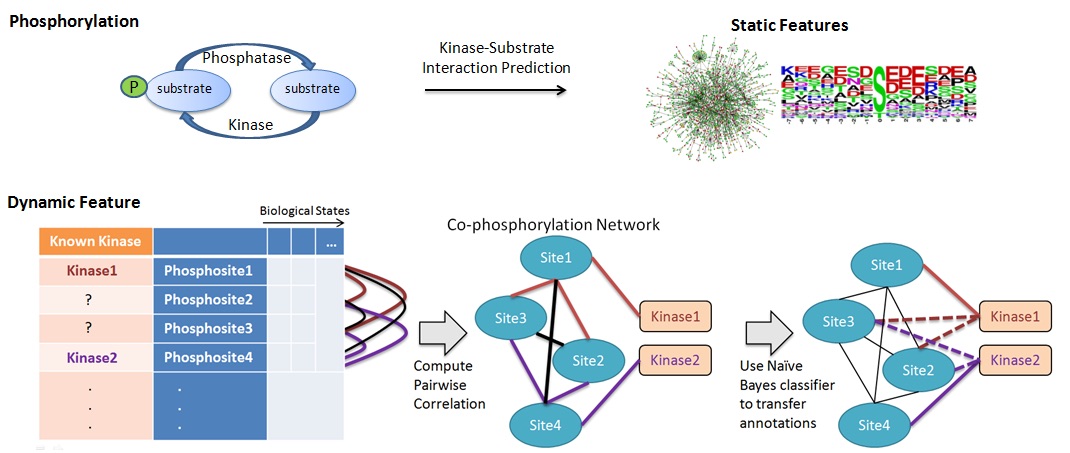
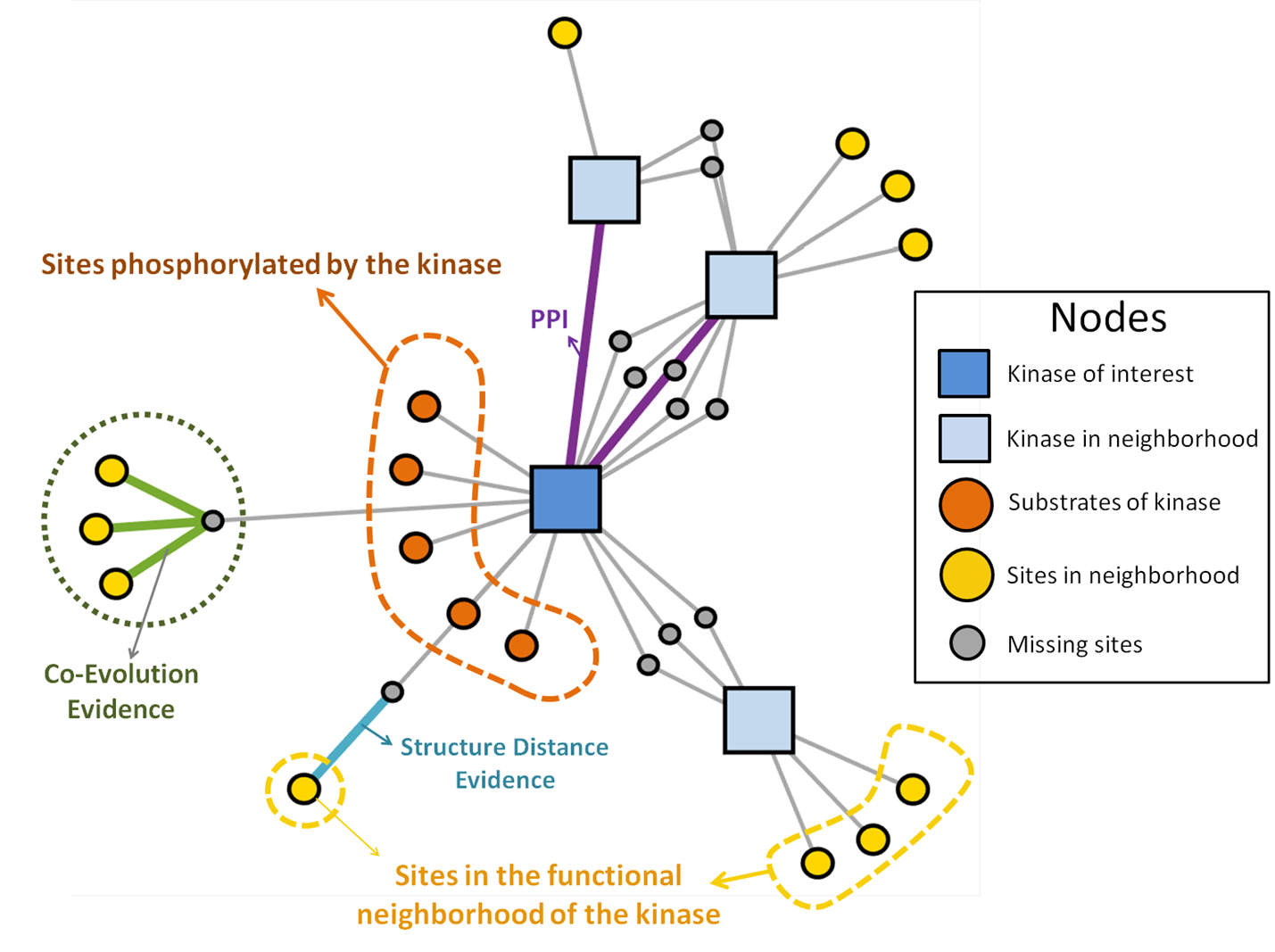
In human cells, attachment of a phosphate to a protein at certain sites can alter the activity and the function of the protein. This mechanism, known as protein phosphorylation, is often used to communicate signals within and between cells. Recent research shows that likely over 70% of human proteins can be phosphorylated. Dysregulation of protein phosphorylation is known to play an important role in many diseases, including cancer, Alzheimer's disease, Parkinson's disease, obesity and diabetes, and fatty liver disease. Indeed, many modern drugs used to treat various cancers target kinases, the enzymes that are responsible for the phosphorylation of proteins. Despite the success of the "genomic revolution" and the importance of protein phosphorylation in human biology, the knowledge on protein phosphorylation in humans is quite limited. To date, thousands of phosphorylation sites on human proteins have been discovered, but the kinases that are responsible for phosphorylating these sites could be identified for less than 5% of these sites. Recognizing the challenges associated with analyzing phospho-proteomic data, we utilize network science to extract patterns of correlation in phosphorylation levels of proteins. By organizing these patterns in "co-phosphorylation networks" and using graph-theoretic algorithms and machine learning, we extract knowledge from these networks, which are then used to develop new biological hypotheses. Besides generating basic biological knowledge such as functional annotation of phospho-proteins, kinases, and phosphatases, we also develop methods to characterize the signaling processes that are affected in cancers and Alzheimer's disease. We collaborate on this project with Mark Chance, Director of the Center for Proteomics and Bioinformatics at CWRU School of Medicine. This project is supported by National Institutes of Health grant R01-LM012980 from the National Library of Medicine. Group members working on this project: Serhan Yılmaz, Tyler Cowman, Filipa Blasco Tavares Pereira Lopes | |
Integration, Compression, and Version Control of Big Networks | |
 | |
|
In many applications, network models are commonly used to represent interactions and higher-level
associations among various entities. Integrated analyses of these interaction and association data
has proven useful in extracting knowledge, generating novel hypotheses, and developing predictive models.
Applications include recommender systems, disease gene prioritization, network de-noising, and tracking temporal
evolution of networks.
Our research seeks to answer a number of fundamental questions that relate to efficient utilization of
large network-structured datasets: - what are (provably) optimal storage schemes for large network structured
databases? how should multiple versions of same/ related datasets be stored? how does one trade-off compression
with query efficiency? and how does one suitably abstract network data so that users can interactively interrogate
them using web-based front-ends?
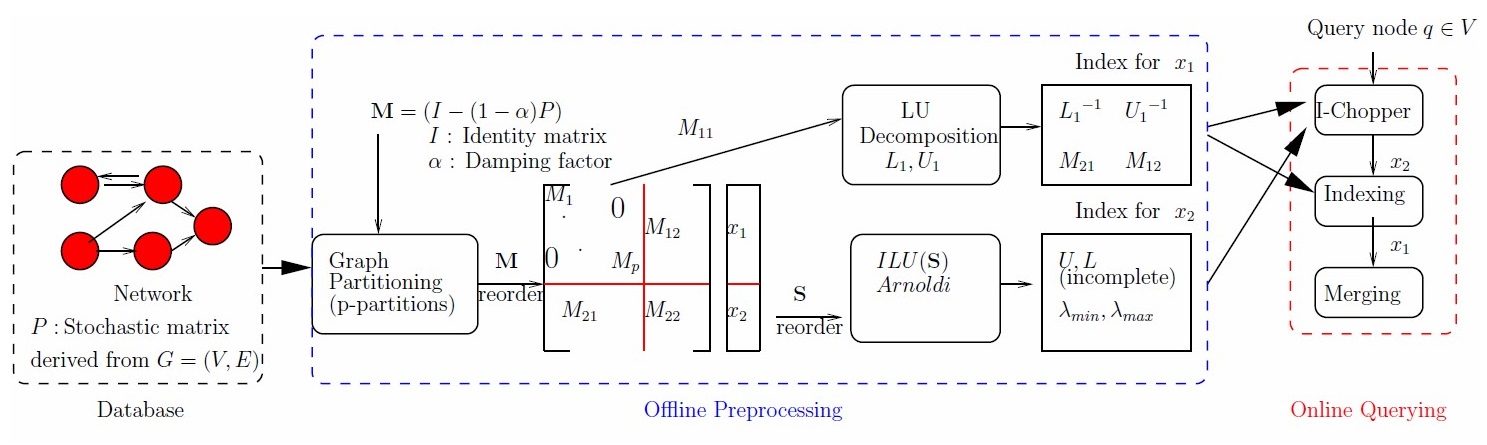
To answer these questions, we develop theoretically grounded and computationally validated storage
schemes, algorithms, and software that enables efficient and effective storage, update, processing,
and querying of big and heterogeneous networks.
This project has been supported by National Institutes of Health grant
U01-CA198941 through the Big Data to Knowledge (BD2K)
program. Group members working on this project: Tyler Cowman, Kaan Yorgancıoğlu, Mengzhen Li |
